Garden Eden with automated Permaculture: Is it possible?
A new way to do automated sustainable farming?

Can one project make a difference? Does code change anything? In a world that increasingly blends virtual and real worlds together, I believe it does. Especially looking at recent events and how their ethical virtual missteps considerably impacted the real world!
In this light, its even more important to come up with ideas and projects that empower people to make better decisions and improve their standard of living. Machine learning (AI, to use the marketing term) has a huge potential to provide us with smarter tools, tools that can make small decisions and present their operators with higher level choices. It’s not like getting a bigger hammer, it’s a hammer that suggests what nails to use. This post is the start of forging such a hammer.
Small scale farming
Farming is one of the areas where the digital “disruption” (read: influence) still is fairly limited, especially on smaller scales. Available solutions are often too expensive or complex to deploy on fields that really should only feed the family and some close friends! In particular the space around permacultures is relying on getting human experts or relying on infographics. If this information was available to computers however, imagine what a robot like farmbot can do with this!
Permacultures
To allow the design and creation of sustainable ecosystems, a wide range of information is required. This information is collected inside the framework that is permaculture. This framework provides principles created, implemented, and expanded by a world wide community of enthusiasts; but it’s origins go back to the 1970’s book “Permaculture One” by Bill Mollison and David Holmgren.
While those principles touch every aspect of living in an “integrated, evolving system of perennial or self-perpetuating plant and animal species useful to man” (cited from Permaculture One), our experiment focusses on the agricultural aspect: creating plant polycultures. To find out more about the definition, origin and anything else head over to https://holmgren.com.au/about-permaculture/!
In essence, followers of these principles divide their living space into zones, each of which can then be refined to reflect a more natural, cooperative, and self-perpetuating approach. Polycultures are only a part of the tools available to shape these zones.
Polycultures
An ecosystem inspired by nature naturally provides a wealth of improvements over regular growing. The plants complement each other and those so called guilds yield benefits such as:
- Fewer or no pestizides required
- Growth nutrients are exchanged between plants (fertilization)
- Larger and more nutritious yields due to better growing conditions
- Improved efficiency
… or in a word: sustainability.
There are a ton of approaches and ways to achieve this, most of which require a lot of experience and humans “looking at the plants” as well as in-depth knowledge of the plants and soil conditions. A daunting task for many newcomers to that space and unavailable to those without experts nearby. Here, we are going to focus on an engineering approach, starting with an MVP 😁 to create valid (and sometimes new) guilds from easily accessible information.
Important influences include:
- Wind
- Sunlight
- Temperature and climate
- Soil properties and type
- Water levels, availability, and rainfall
- Insects, polination, flowering and harvesting times
Information about the plans themselves is hard to come by, but we were lucky that the people over at https://permacultureplantdata.com collected and structured data of about 2000 plants. This will be the data we are working with, and it includes:
- Root depth and spread. Clearly roots needs water but they can only work well if they don’t compete for the same reservoir. These need to be as far apart as possible.
- Flowering and harvest times. The most efficient garden generates yield year-round without “empty” months. Yet there can’t be too many plants flowering at the same time since this adds stress to their companions!
- Soil types and PH values. Plants won’t grow large in suboptimal soil conditions and while they can be changed (e.g. with fertilization), a guild has to overlap on soil properties as much as possible.
- USDA hardiness zones. These are zones defined by the US Department of Agriculture that classify climate, elevation and soil properties. This is a great proxy for these values and they need to overlap!
Goals, and how to get there
Based on the ideas of a colleague of mine, we - a group of seven Microsoft employees - took on the challenge of starting this project during a one-week internal hackathon. The goal was to create a system that is able to generate an optimized polyculture for home use. Since then, our efforts have come a long way and it has become - mathematically speaking - a combinatorics problem, where we want to maximize yield while satisfying constraints for each plant as well as possible.
However, these constraints change with every plant that is added to the mix, letting us determine the “value” of a mix of plants only after the combinations are known. Effectively it’s necessesary to come up with a combination, then start to evaluate. By comparing these evaluations (e.g. via a score) we could then rank and find the best ones.
However by using a brute force approach (enumerate all possible combinations of a given set), the runtime will grow exponentially - at O(n!), which means that it might take decades or longer to calculate!
So to generate this result set quickly (in a reasonable time), a more structured approach is required. One such approach can be framing the problem as an optimization problem in order to apply a metaheuristic approach! In our case, genetic algorithms with their permutating power seem to be well suited.
Guild Wars
For our approach an individual (i.e. a potential solution to the problem) is essentially a guild - a set of plants growing together, which will be chosen at random from all the available plants.
Next, we require a fitness function to evaluate the fitness of each those individuals! This is the tricky part since we want to create guilds to score highly when they work out in the real world. As mentioned above we only a have a few features to go on, so - for now - the fitness function will be evaluated with the main plant:
def fitness(individual, main_plant_id):
if len(individual) == 0:
return (-10000,)
main_plant = plants_by_id[main_plant_id] # resolve the plant id to all plant data
score = 0
# initialize the score with penalties/bonusses for known companions
if "_companions" in main_plant:
compatible_ids = [plants_by_commonname[p["Companion"]]["id"]
for p in main_plant["_companions"] if p["Compatible"] == "Yes"]
incompatible_ids = [plants_by_commonname[p["Companion"]]["id"]
for p in main_plant["_companions"] if p["Compatible"] == "No"]
for i in individual:
if i in compatible_ids: score += 1000
if i in incompatible_ids: score -= 1000
# add the main plant to the mix for further scoring
plants = [main_plant] + [plants_by_id[i] for i in individual]
# standard deviation on roots
roots = [root_depth(p) for p in plants]
root_depth_spread = np.std(roots)
# overlap in soil requirements
soil_intersect = reduce(lambda x, y: x.intersection(y),
[enum_set(ph(p)) for p in plants], set())
# overlap in USDA zones
usda_intersect = reduce(lambda x, y: x.intersection(y),
[enum_set(usda(p)) for p in plants], set())
# common memberships
membership_intersect = reduce(lambda x, y: x.union(y),
[memberships(p) for p in plants],
set()).intersection(memberships(main_plant))
# common uses (max 10)
uses = set()
for plant in individual:
u = [u["Use"] for u in plant["_uses"]] if "_uses" in plant else []
uses.update(u)
# finalize scoring
score += 10 - len(uses)
score += len(soil_intersect)
score += len(membership_intersect) * 100
score += len(usda_intersect)
score *= (root_depth_spread if not np.isnan(root_depth_spread) else 0)
# make score independent of guild size
return (score / len(individual),)
In short, the fitness function prefers high overlap in soil ph values, USDA zones, known guild membership and use category, as well as a known compatibility (while punishing known incompatibility heavily). Currently the only thing to prefer a large spread (i.e. a high standard deviation) in is the root depth. As a first draft this fitness function serves its purpose, but it will likely get a lot more complex over time.
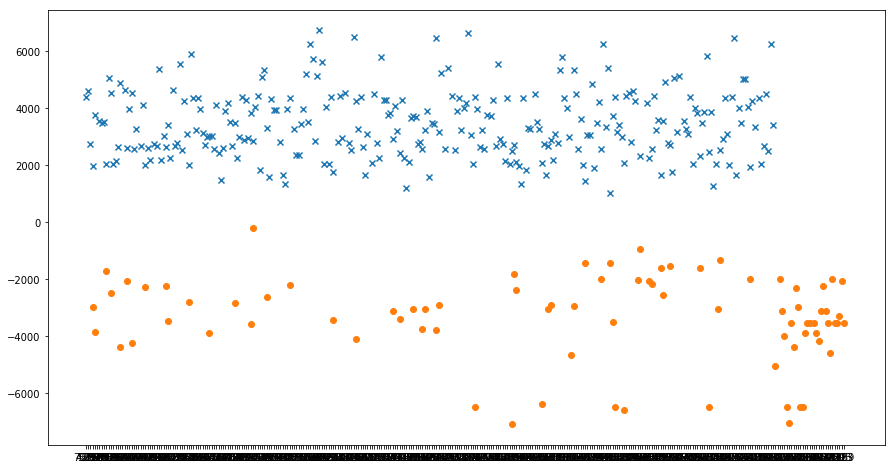
Fitness of known compatible (x) and incompatible (o) plants
As expected the function separates known compatibilites from incompatibilities.
Mutation and Crossover
Mutation and recombination/crossover are two very important operators to introduce some sort of randomness into the population. This can be very sophisticated, but for now they either add another plant to the guild with a 50% chance (mutation) or using something called a two point crossover.
Selection
Simple - only the best µ (MU in the code below) parents are selected to create the next population. This makes sense as a starting point but might change in the future when there’s a deeper insight into the problem.
Let the science begin!
The framework we used to work with genetic algorithms is called DEAP, which is nicely documented and comes with a lot of prebuilt tools and algorithms.
Its basis is a toolbox class that registers the individual steps and calls them as appropriate. Since we are working with a custom population (a list of string ids), the initial population has to be created using a specialized function (see below). To get to know all the ins and outs of the toolbox etc, please check out their excellent tutorials. This is our first shot at solving this, so we started with a randomized set of plants!
creator.create("FitnessMax", base.Fitness, weights=(1.0,)) # Maximization, change weights to -1.0 to minimize
creator.create("Individual", list, fitness=creator.FitnessMax)
toolbox = base.Toolbox()
# Population creation functions
toolbox.register("individual_guess", initIndividual, creator.Individual)
toolbox.register("population_guess", randomPopulation, list, toolbox.individual_guess, main_plant_id)
# GA operators
toolbox.register("evaluate", fitness)
toolbox.register("mate", tools.cxTwoPoint)
toolbox.register("mutate", mutSet)
toolbox.register("select", tools.selBest)
# GA parameters
NGEN = 15 # number of generations
MU = 100 # number of individuals to select each round
LAMBDA = 1000 # population size of each round
CXPB = 0.5 # crossover percentage
MUTPB = 0.5 # mutation percentage
pop = toolbox.population_guess()
hof = tools.ParetoFront() # hall of fame to keep the absolute best individuals
# statistics output
stats = tools.Statistics(lambda ind: ind.fitness.values)
stats.register("avg", np.mean, axis=0)
stats.register("std", np.std, axis=0)
stats.register("min", np.min, axis=0)
stats.register("max", np.max, axis=0)
algorithms.eaMuPlusLambda(pop, toolbox, MU, LAMBDA, CXPB, MUTPB, NGEN, stats, halloffame=hof)
Each generation produces nice fitness statistics and in order to see if the algorithm converges properly standard deviation has to go down, while the average should go up 🖖:
gen nevals avg std min max
0 100 [1134.57323392] [10255.55109512] [-92857.64144998] [25895.75306028]
1 1000 [12593.19622808] [12802.6623178] [5873.33588997] [120468.65434407]
2 1000 [50537.4606231] [31176.55527062] [25895.75306028] [224921.81501051]
3 1000 [138444.15673245] [44540.70143468] [98907.68243759] [325421.74504245]
4 1000 [272949.4089385] [45394.31901551] [224390.73227114] [369011.82987117]
5 1000 [366902.72634401] [44232.56633966] [325870.60453851] [454566.55372268]
6 1000 [457777.01465387] [42910.76824181] [425915.83348434] [623003.72087292]
7 1000 [566856.99261163] [49954.55599536] [526699.08958865] [722884.90679263]
8 1000 [664096.17029895] [45880.71422853] [624681.71366634] [724121.04313177]
9 1000 [724327.28180196] [4460.43376957] [722884.90679263] [746783.22084422]
10 1000 [728910.31333747] [9025.99582626] [724121.04313177] [746783.22084422]
11 1000 [747677.8868938] [8901.81479744] [746783.22084422] [836249.82580286]
12 1000 [756624.54738967] [27993.18428005] [746783.22084422] [836249.82580286]
13 1000 [833629.86000009] [15286.39903347] [746783.22084422] [842653.06040163]
14 1000 [837044.78445508] [2083.14210111] [836249.82580286] [843809.60703609]
15 1000 [843002.72909254] [2057.02973603] [842653.06040163] [863168.44631953]
Plotting these numbers shows great convergence over these 15 generations and we could even try and run a bit longer to get better solutions.
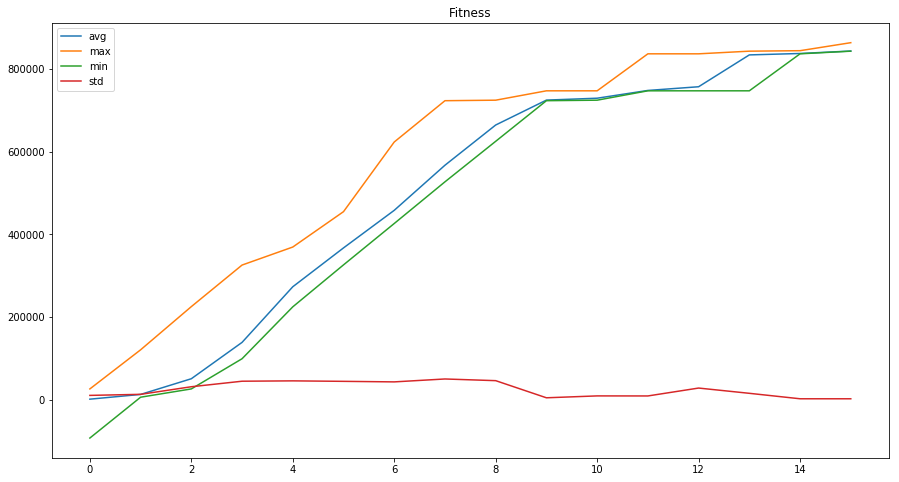
So far the experiment looks very promising, we will continue working on validating the results in order to achieve real usefulness 😊.
To our future selfs
The current results are already quite promising and show a good direction already. We have several next steps planned:
- Improve data quality (normalize verbose fields, etc.)
- Improve the scoring to reflect real world usefulness
- Generally improve the algorithm and validate results 😊
As this is a really hard real-world problem, we also needed to reduce complexity and thus the decision was made to simplify some things:
- Plant placement and size are left out completely
- Soil and climate data is mostly abbreviated by using the USDA hardiness zones
- Growth time is left out as a factor
- At most one target plant is required (i.e. I have an apple tree, what goes with it?)
- Guilds are optimized for a single purpose (e.g. food only)
As we continue the project there will be more of those that come into consideration, but so far we think we are on a promising path. There will be more blog posts covering this topic, so look out for those!
Until then there will be more Rust content, and a new format! To stay updated on those exciting developments, subscribe via RSS (like feedly) and follow me on twitter.







